|
2/17/2024 0 Comments Promoter consensus sequenceApplying CFPS for the rapid discovery of bioactive molecules is particularly interesting as it allows transforming functional metagenomics to host-independent systems in high-throughput platforms. In addition to diversifying the areas of application, the main focus of CFPS has been increasing the quantity and quality of synthesized protein. Cell-free protein synthesis (CFPS) has been effectively used in a variety of applications including the production of difficult-to-express proteins for reasons of toxicity or solubility. To bypass the use of cell host for expression of metagenomic DNAs, a previous study has demonstrated the use of a recombinant bacterial transcription system to generate mRNA from environmental DNAs 8. The main setback of functional metagenomics is associated with the host systems where transcription of foreign DNAs is greatly biased which subsequently result in low hit rate with limited diversity 4, 5, 6, 7. Due to all these, there is an increasing non-linearity between sequence mining and getting active corresponding protein which requires a different innovative approach to address. In addition, a series of subcloning steps for the positive hits have to be performed to reach the target genes. It also requires a lengthy and laborious process of generating a large number of clone libraries from environmental DNAs. Functional metagenomics, despite its potential to yield truly novel enzymes, has suffered serious challenges in protein expression. Nonetheless, the task of functional identification of biomolecules heavily relies on the successful expression and biochemical verification of target genes. 
The exponential increase of sequence data and rapidly evolving smart bioinformatics tools are making in silico predictions from these resources more successful. It is a powerful tool that can reach the untapped vast majority of microbial resources to answer questions about diversity and function 1, 2, 3. Metagenomics has played a vital role in the discovery of novel biomolecules in the last few decades.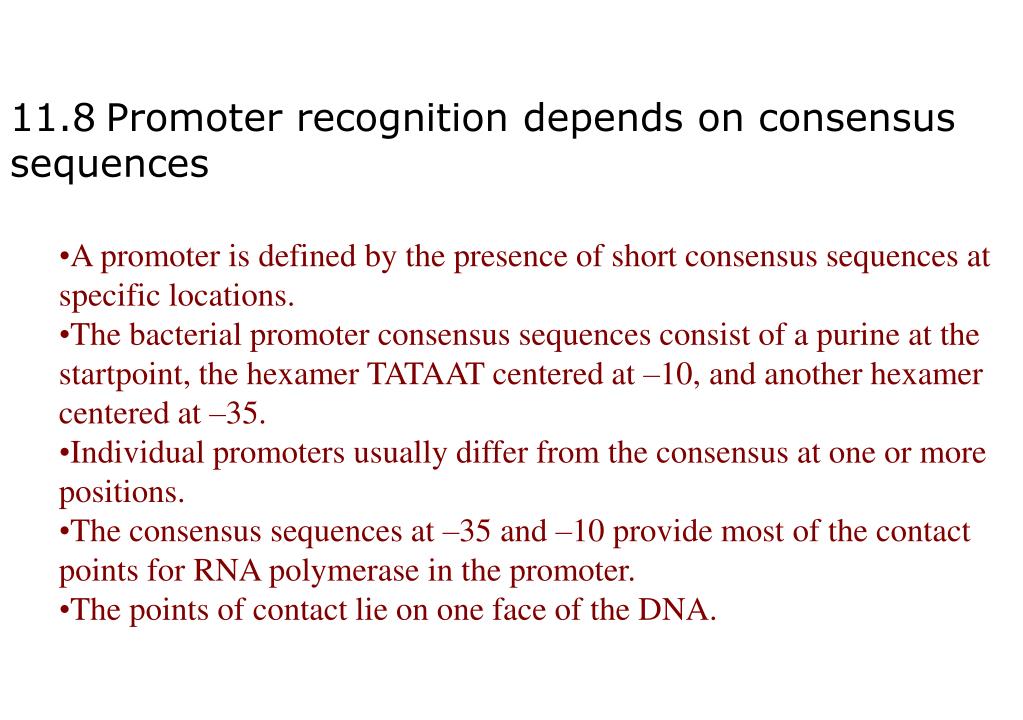 
Here, we have successfully identified and verified 12 enzymes acting on bis(2-hydroxyethyl) terephthalate (BHET) in a completely clone-free approach and proposed an in vitro high-throughput metagenomic screening method. 
Using EM1 RNAP and a translation-competent Escherichia coli extract, we have developed an efficient medium-throughput pipeline and protocol allowing the expression of metagenome-derived genes and the production of proteins in cell-free system is sufficient for the initial testing of the predicted activities. EM1 RNAP and its promoter sequence are distantly related to T7 RNA polymerase. We here report the initial characterization of novel single-subunit bacteriophage RNA polymerase, EM1 RNAP, identified from a metagenome data set obtained from an elephant dung microbiome. However, function-based metagenome searches are often limited by the time-consuming expression of the active proteins in various heterologous host systems. The mining of genomes from non-cultivated microorganisms using metagenomics is a powerful tool to discover novel proteins and other valuable biomolecules.
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
